What can we learn from previous SARS outbreaks
- March 27, 2020
Every problem has already been solved by someone. That’s why efficient innovation starts with researching existing knowledge across industries and domains. When learning from existing solutions, the wheel is not reinvented, and the risk factor related to innovation is reduced subsequently. And we save precious time. Creax wondered to what extent this philosophy is being applied for dealing with the corona crisis.
The COVID-19 (coronavirus disease 2019) pandemic we are facing right now is caused by a coronavirus which is related (about 79% genetic similarity) [1] to the virus that caused the SARS (severe acute respiratory syndrome) epidemy in the 2000s. The virus belongs to the species SARS-related coronavirus (SARS-CoV) and has been named SARS-CoV-2 [2]. Thanks to the similarity with the SARS coronavirus (SARS-CoV or SARS-CoV-1), and other SARS-related coronaviruses (SARSr-CoVs), previous scientific research on these viruses could provide valuable insights for the COVID-19 pandemic. Therefore, Creax analyzed a pool of peer-reviewed scientific documents mentioning SARS-CoV-1 or SARSr-CoVs and SARS-CoV-2. The goal is to reveal possible patterns in the responses of the scientific and industrial community to these outbreaks, and to extract potential implementations of previous learning into the COVID-19 context
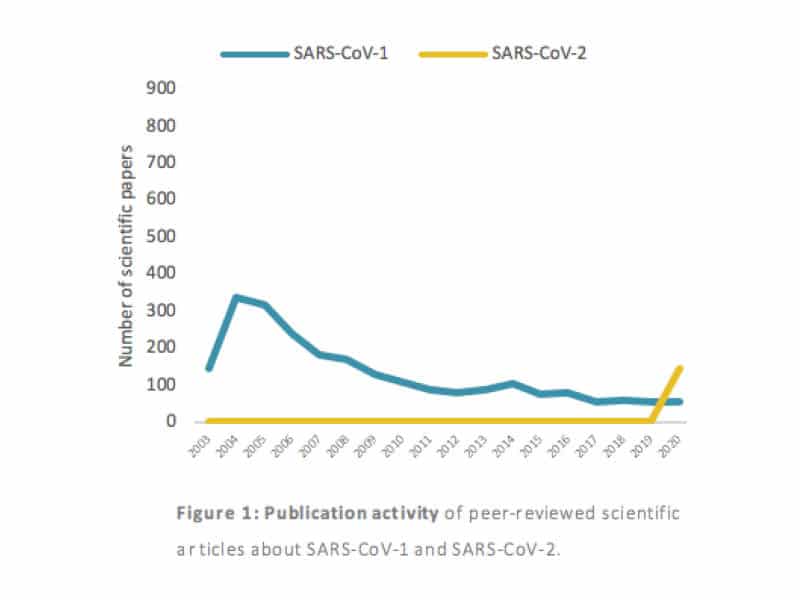
Figure 1 shows the number of papers published between 2003, the year of the SARS-CoV-1 outbreak, and today. While the blue track is giving the trend for papers specifically mentioning SARS-CoV(-1), the yellow track relates to papers focusing on SARS-CoV-2. A steep growth of SARS-CoV-2 publications in 2020 is clearly noticeable. Only peer-reviewed publications are included in this pool. In fast-evolving situations like this, it is recommended to capture early-stage findings in scientific research. Therefore, a new pool of papers was built for which pre-printed (thus not peer-reviewed) manuscripts were included.
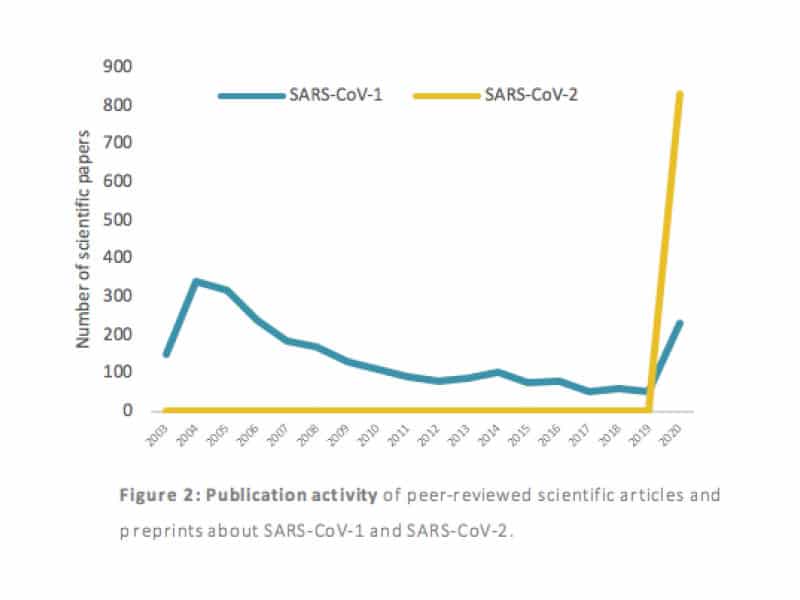
Figure 2 shows a similar trend line for SARS-CoV-1 papers but the yellow line, showing a drastic increase in 2020, reveals the big effort the scientific community is making to gather insights on COVID-19. And it is on-going. Between March 23rd and March 25th, around 65 new COVID-19 SARS-CoV-2 preprints were published, bringing its number to 765.
The scientific community is responding very fast to the COVID-19 pandemic.
To get an overview of the topics discussed in these scientific publications, the document pool was analyzed by the Creax team, supported by its in-house developed (con)text-mining platform Mynd. Mynd offers the possibility to get profound insights into a big collection of text documents from various sources. The software allows us to create an unbiased, detailed, and at the same time exhaustive view about a scientific or technological domain. This results in learnings about trends, emerging technology developments, important applications, white space opportunities, etc… It is applicable for both experts in their fields as for people/companies that want to discover a domain without having any knowledge or experience.
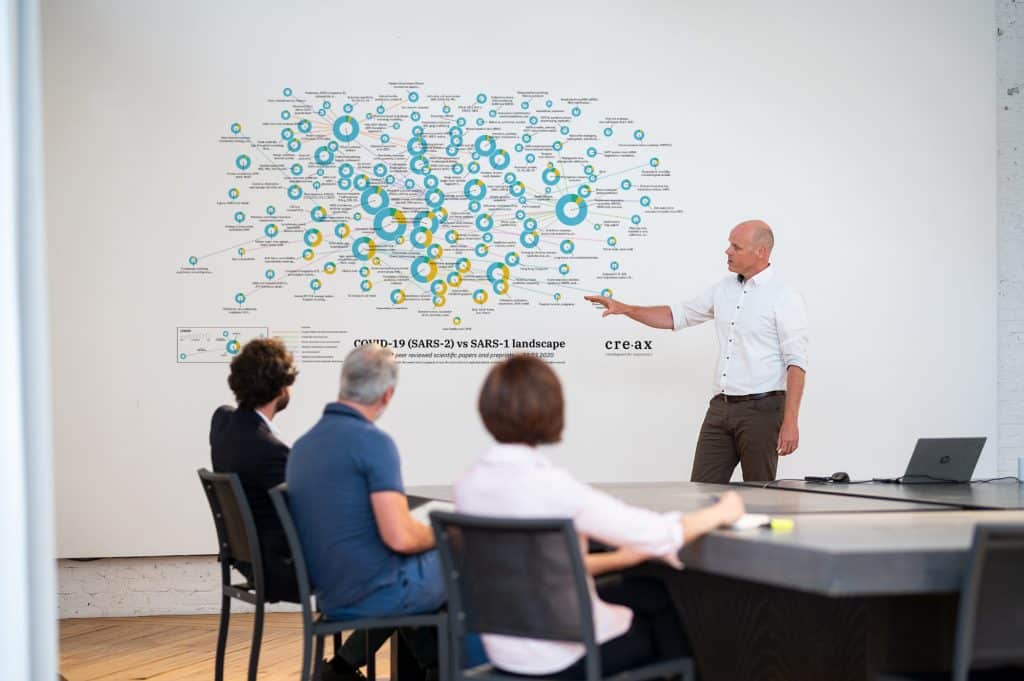
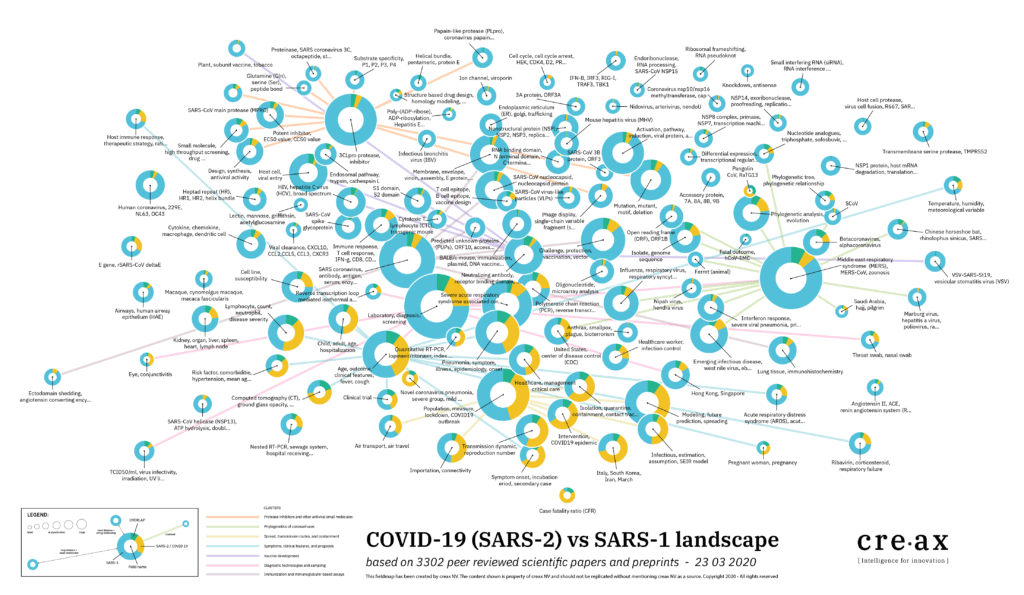
A first outcome of the Mynd analysis is a field map (figure 3). This field map visualizes the clustering of documents that discuss similar topics. Each circle represents a specific field of interest. The annotation of each circle describes the keyword(s) relevant to this field. What can we learn from this graph?
The size of the circle indicates how many articles are linked to the field. The coloring of the circle is linked to papers discussing SARS-CoV-1 (cyan) and SARS-Cov-2 (yellow). Interesting are the green overlaps. These indicate an activity field in which learnings from the previous SARS outbreak are discussed in function of current scientific research about COVID-19. The proximity of the bubbles indicates how closely related topics are. As such we can allocate different groups to different bigger domains of interests.
In the lower part of the map, a higher percentage of papers about SARS-CoV-2 is seen. Almost all these papers discuss the epidemiology, disease control, and diagnosis of COVID-19. This is not surprising as in response to a fast-evolving pandemic, like COVID-19, priority focus is laid on monitoring the infection rate and spread, methodologies to reduce the impact of the infection (e.g. flatten the curve), and diagnosing the disease through a fast, sensitive and reliable test.
First responses to the COVID-19 pandemic are focusing on 1) monitoring the spread of the virus, 2) strategies to reduce the spread of the virus, and 3) development of diagnostic tests.
Researchers are clearly looking to learn from previous virus outbreaks. As an example, in the right part of the field map a big cluster about “Middle East respiratory syndrome (MERS), MERS-CoV” is detected. Close to this field, fields about ‘beta and alpha-coronavirus’ and ‘phylogenetic evolution’ are found. Hence, scientific papers connecting these fields of interest focus on the genetic relationship between viruses and research groups learn from that.
Researchers also rely on established strategies. Amongst others, a big field of interest in the left corner of the field map is about 3CLpro protease, which is a family of enzymes that plays a role in the replication of coronaviruses.[3] The field does not seem to be populated by a lot of articles about SARS-CoV-2. However, because of the low threshold of the Mynd platform, a preprint paper by Stoermer et al. was found that states that 3CLpro protease of SARS-CoV-2 is for 96% identical to the 3CLpro protease of SARS-CoV-1.[4] Inhibitors previously developed for the SARS-Cov-1 variant potentially could be used to inhibit the reproduction of SARS-CoV-2 and thus in the treatment of COVID-19. Natural polyphenols have been mentioned in this context as suspected to be more effective than e.g. nelfinavir.[5]
Researchers learn from previous virus outbreaks and rely on established strategies to treat for COVID-19 that are sometimes based on natural products.
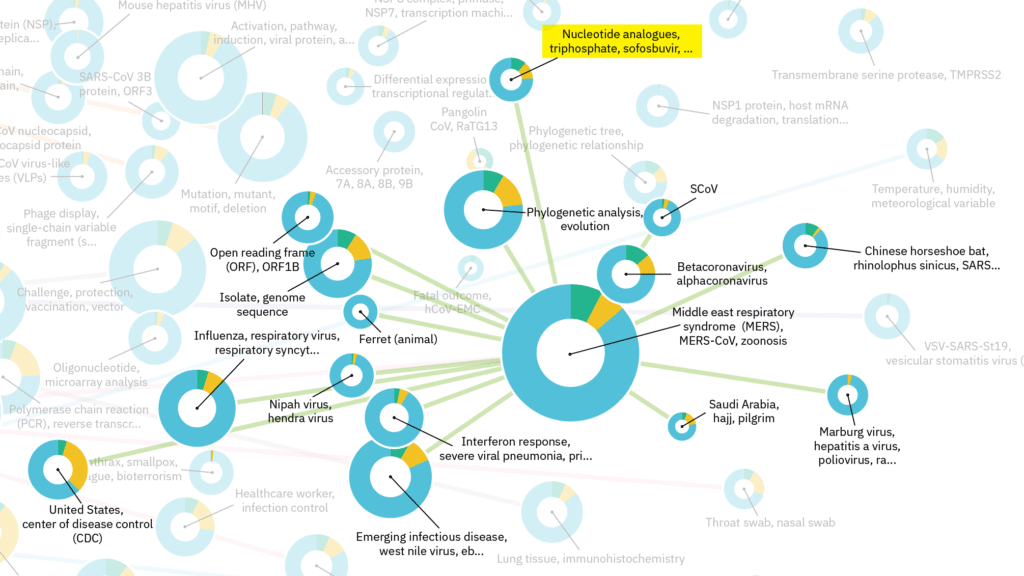
As informative the individual fields are, novel solutions can sometimes be found out domain through associations between fields. As referred to at the beginning of this blog, researchers do not always need to reinvent the wheel. Let’s take the ‘MERS’ cluster as an example (figure 4). This cluster links fields about virus families, their originating animals (e.g. pangolin, horseshoe bat, ferret, etc.) and regions, and therapeutic strategies. One interesting associated field is the one marked in yellow in figure 4 (‘Nucleotide analogues’). In this field, a preliminary research activity was found about the use of nucleotide analogues to inhibit RNA-dependent RNA polymerase (RdRp), which is also relevant for SARS-CoV-2. These broad-spectrum nucleotide analogues have shown to be effective for SARS-CoV-1 before and it has been suggested to develop a therapeutic for SARS-CoV-2 based on these compounds[6]. A potential nucleotide analogue drug in the battle against COVID-19 might be based on the FDA-approved hepatitis C drug EPCLUSA (Sofosbuvir/Velpatasvir) developed by the pharmaceutical company Gilead Sciences.
Despite not being an expert in each individual scientific field, Creax is able to discover and quantify in an unbiased way the scientific dynamics evolving since the COVID-19 outbreak, thanks to its in-house developed AI platform Mynd. As such we have revealed that learnings about monitoring, limiting, diagnosis or therapies for SARS-CoV-2 are reused or used as inspiration in the current context.
The field maps shown in this blog are only the starting point for deep-dive research. The Mynd platform allows us to filter out specific topics and categorize them accordingly, for example as therapeutic compounds, biological targets, target groups, etc. The combination of these specific types of information is a catalyst for efficient research roadmaps.
The Creax team is available to answer your specific interests in this research and discuss potential requests for deep-dive analysis of interest areas. Please find our contact details below.
References
[1] Figure 1 Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 2020; 395:565–74.
[2] Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol 2020; 5.
[3] Kim Y, Lovell S, Tiew K-C, Mandadapu SR, Alliston KR, Battaile KP, Groutas WC, Chang K-O. Broad-Spectrum Antivirals against 3C or 3C-Like Proteases of Picornaviruses, Noroviruses, and Coronaviruses. J Virol 2012; 86:11754–62.
[4] Stoermer M. Homology Models of Coronavirus 2019-nCoV 3CLpro Protease. ChemRxiv 2020;
[5] Adem S, Eyupoglu V, Sarfraz I, Rasul A, Ali M. Identification of potent COVID-19 main protease (Mpro) inhibitors from natural polyphenols: An in silico strategy unveils a hope against CORONA. 2020; :1–29.
[6] Ju J, Kumar S, Li X, Jockusch S, Russo JJ. Nucleotide Analogues as Inhibitors of Viral Polymerases. bioRxiv 2020; 1:2020.01.30.927574.
Download this paper

Related blogs

Innovation in times of crisis
The corona crisis is an unforeseen challenge that we need to solve as quickly as possible. And a lot is already happening. The confusing and uncertain situation highlights how crucial innovation really is.

Working from home – experiences from Creax
The Corona virus forces us to work together in a new and isolated way. Thanks to the internet, we can work from a remote location, but there are some hurdles along the way. Here are some of our experiences and tips for working from home.

Stop Corona! Download these posters
In times of crisis, what people need most is clear and correct information. We made posters that you can download with the two main messages to stop Corona: keep your distance + wash your hands